
HTRF Human Total FOXM1 Detection Kit, 10,000 assay points
This HTRF kit allows the cell-based quantitative detection of Total FOXM1.
| Feature | Specification |
|---|---|
| Application | Cell Signaling |
| Sample Volume | 16 µL |
This HTRF kit allows the cell-based quantitative detection of Total FOXM1.


Loading...
Product information
Overview
FOXM1 (Forkhead box M1) is a transcription factor expressed throughout the cell cycle, rising in the late G1-phase, reaching its apex in the S-phase, and remaining in the G2/M and late M-phases. It stimulates the expression of several genes involved in cell cycle progression, and plays a significant role in promoting cell division and proliferation. In cancer, FOXM1 is a master regulator of tumor metastasis. It promotes the expression of genes involved in cell migration, invasion, and metastatic spread.
HTRF assays offer many advantages over other technologies:
- Homogeneous add-and-read format
- No wash steps
- Low background
- Straightforward miniaturization from 96- or 384-well microplates to high density assay formats such as 384-well low volume and 1536-well plates
- Stable signal, providing flexibility in readout time and assay size
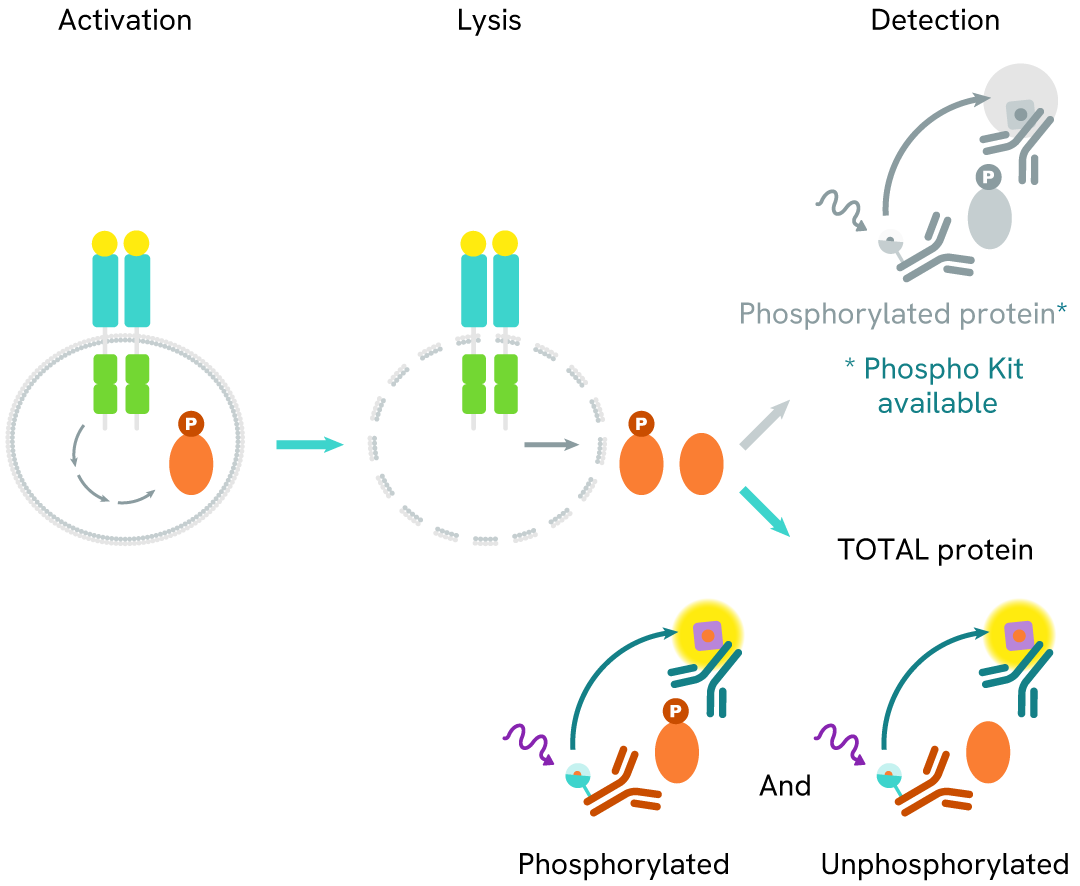
How it works
Total FOXM1 assay principle
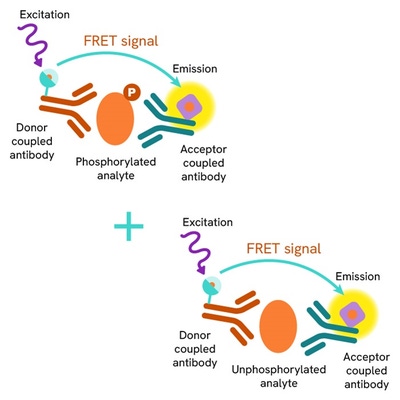
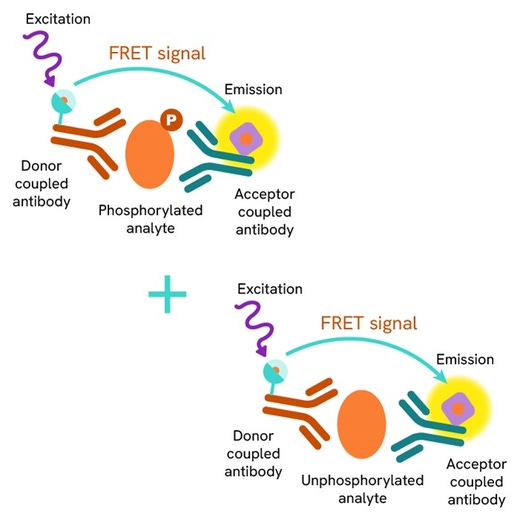
The Total FOXM1 assay measures FOXM1 levels in cells. Unlike Western Blot, the assay is entirely plate-based and does not require gels, electrophoresis, or transfer. The assay uses 2 antibodies, one labeled with a donor fluorophore and the other with an acceptor. Both antibodies are highly specific for a distinct epitope on the protein.
In the presence of FOXM1 this enables an immune-complex formation involving both labeled antibodies. It brings the donor fluorophore into close proximity to the acceptor, thereby generating a FRET signal. Its intensity is directly proportional to the concentration of total protein present in the sample and provides a means of assessing the protein's phosphorylation state under a no-wash assay format.

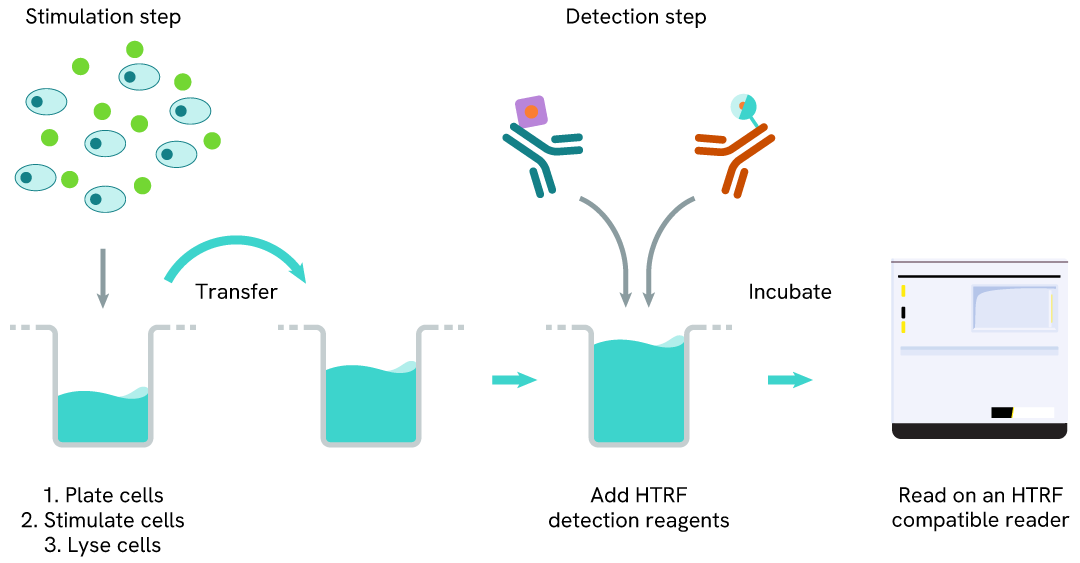
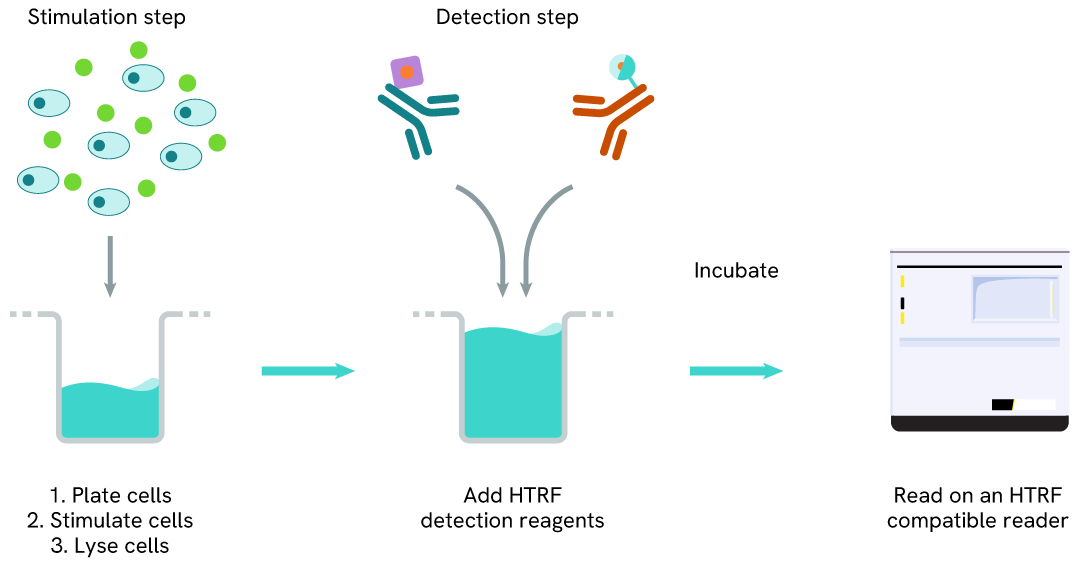
Total FOXM1 two-plate assay protocol
The two-plate protocol involves culturing cells in a 96-well plate before lysis, then transferring lysates into a 384-well low volume detection plate before the addition of Total FOXM1 HTRF detection reagents. This protocol allows the cells' viability and confluence to be monitored.
Total FOXM1 one-plate assay protocol
Detection of Total FOXM1 with HTRF reagents can be performed in a single plate used for culturing, stimulation, and lysis. No washing steps are required. This HTS designed protocol allows miniaturization while maintaining robust HTRF quality.
Assay validation
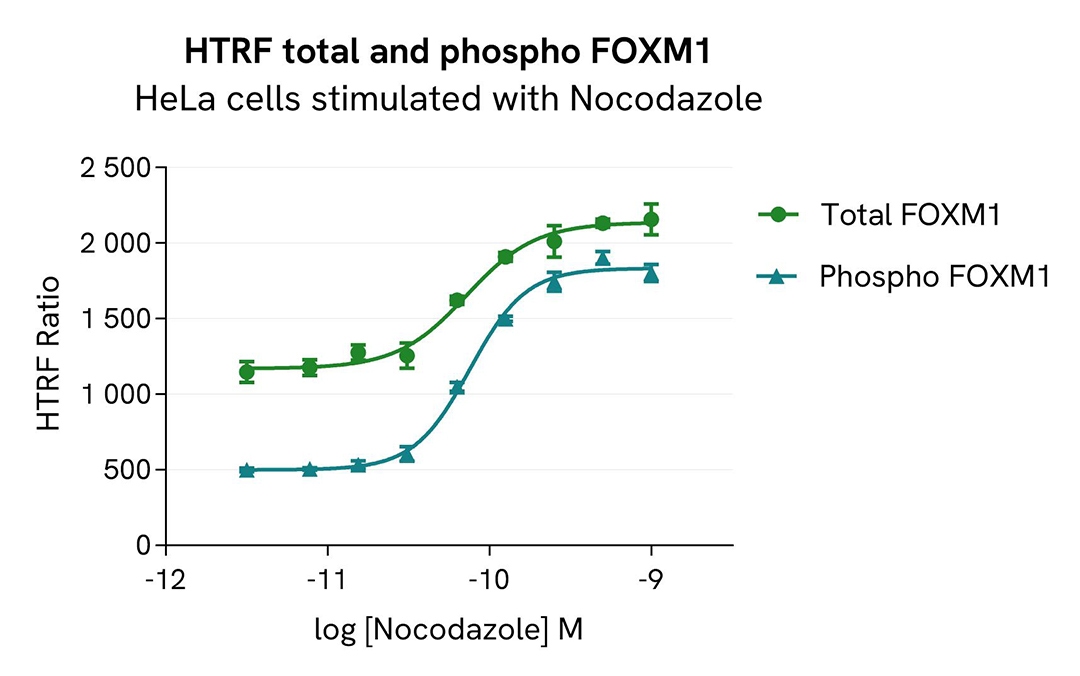
Total FOXM1 modulation using Nocodazole
HeLa cells were seeded in a 96-well culture plate (100,000 cells/well) in complete culture medium and incubated for 24 hours at 37°C, 5% CO2. Cells were then treated for 20 hours with increasing concentrations of Nocodazole.
After cell lysis, 16 µL of lysates were transferred into a 384-well low volume white microplate, and 4 µL of the HTRF Total FOXM1 or Phospho-FOXM1 (Thr600) detection antibodies were added. The HTRF signal was recorded after 4 hours of incubation.
As expected, Nocodazole triggered a dose-dependent increase in total and phosphorylated FOXM1 at Thr600.
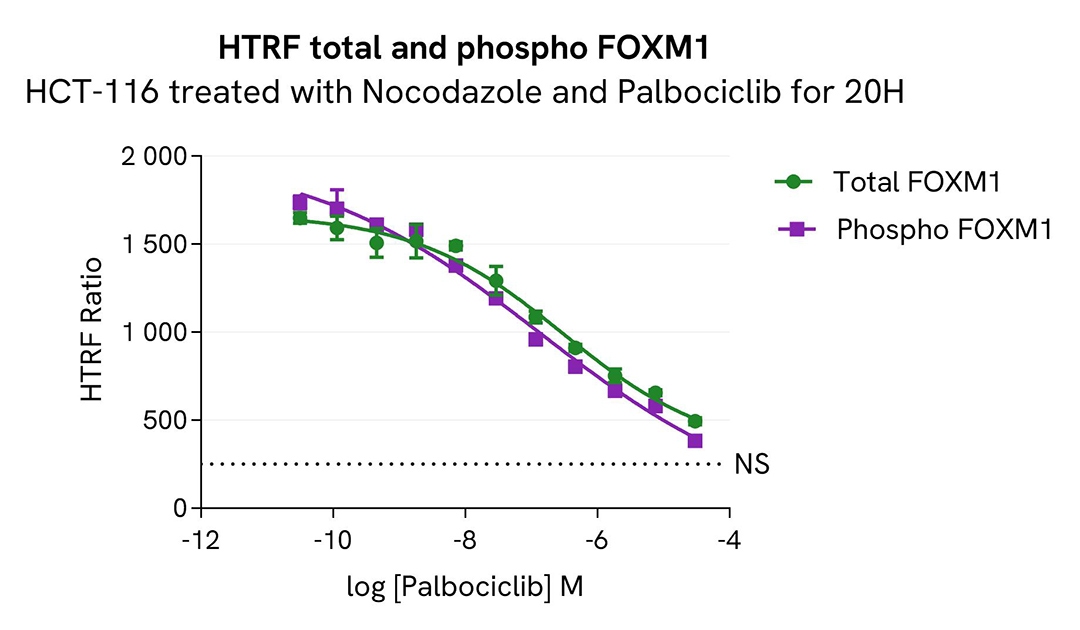
Pharmacological modulation of Total FOXM1 using Palbociclib
HCT-116 cells were seeded in a 96-well culture plate (100,000 cells/well) in complete culture medium and incubated for 24 hours at 37°C, 5% CO2. Cells were then co-treated for 20 hours with increasing concentrations of Palbociclib and 0.5µM of Nocodazole.
After cell lysis, 16 µL of lysates were transferred into a 384-well low volume white microplate, and 4 µL of the HTRF Total FOXM1 or Phospho-FOXM1 (Thr600) detection antibodies were added. The HTRF signal was recorded after 4 hours of incubation.
As expected, Nocodazole with Palbociclib (CDK4/6 inhibitor) treatment induced a decrease in both total and phospho- FOXM1 (Thr600) to the basal level.
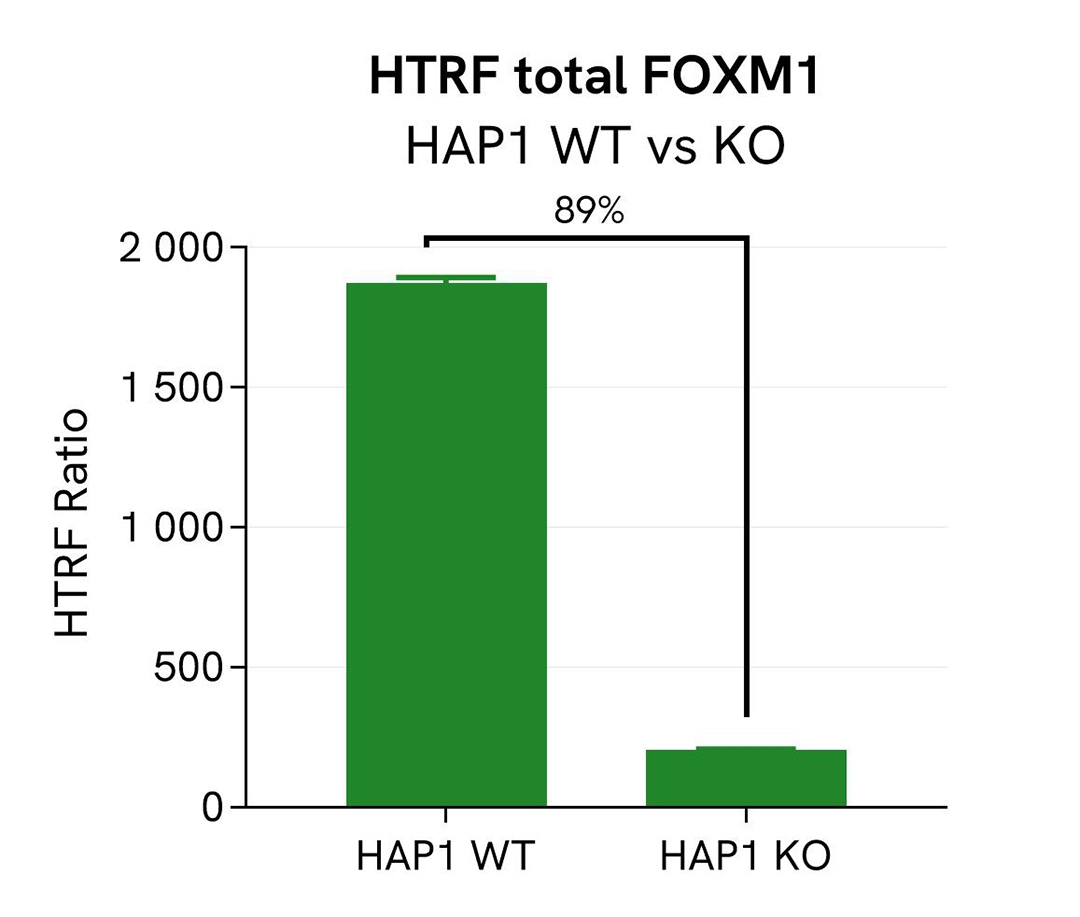
Validation of the specificity of Total FOXM1 using HAP1 cells knocked out for FOXM1
Total FOXM1 protein levels were assessed with the HTRF Human Total FOXM1 kit in HAP1 cells (WT) and a HAP1 cell line Knocked-Out for FOXM1. The cell density was optimized beforehand to ensure HTRF detection within the dynamic range of the kit (data not shown).
The cells were cultured in a 96-well plate (100,000 cells/well) for 24 hours at 37°C, 5% CO2. They were then lysed with 50 µL of supplemented lysis buffer #1 (1X) for 30 minutes at RT under gentle shaking, and 16 µL of cell lysate were transferred into a low volume white microplate, followed by 4 µL of premixed Total FOXM1 detection reagents. The HTRF signal was recorded after 4 hours of incubation at RT.
In HAP1 KO FOXM1 cells, the HTRF signal decreased by 89%, indicating a significant FOXM1 gene silencing, whereas the FOXM1 level was well detected in the other cell line, as expected. This demonstrates that the HTRF Total FOXM1 assay is specific for FOXM1 detection.
Catalog cell line references (Horizon Discovery): HAP1 Wt #C631; HAP1 KO FOXM1 #HZGHC000835c006
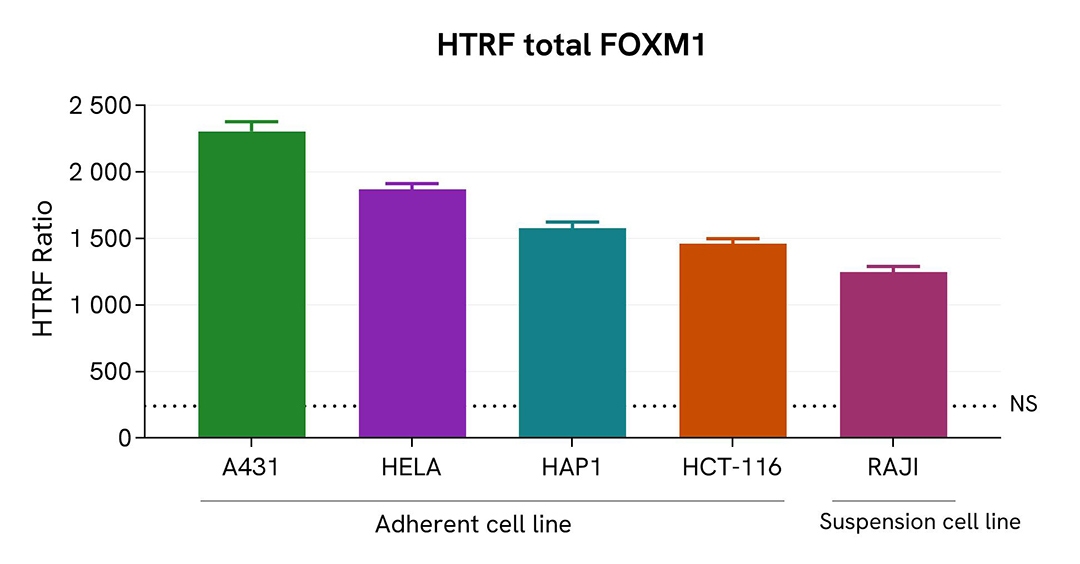
Assessment of Total FOXM1 level in various cell lines
Adherent (A431, HeLa, HAP1, & HCT-116) and suspension (RAJI) human cells were seeded at 100,000 cells/well in a 96-well microplate. After 24h of incubation, the cells were lysed for 30 minutes with supplemented lysis buffer #1, following the protocol for adherent or suspended cells at RT under gentle shaking.
16 µL of lysate were transferred into a 384-well low volume white microplate before the addition of 4 µL of the HTRF Total FOXM1 detection reagents. The HTRF signal was recorded after 4 hours of incubation.
The HTRF Total FOXM1 assay efficiently detected Total FOXM1 in various cellular models expressing different levels of the protein.
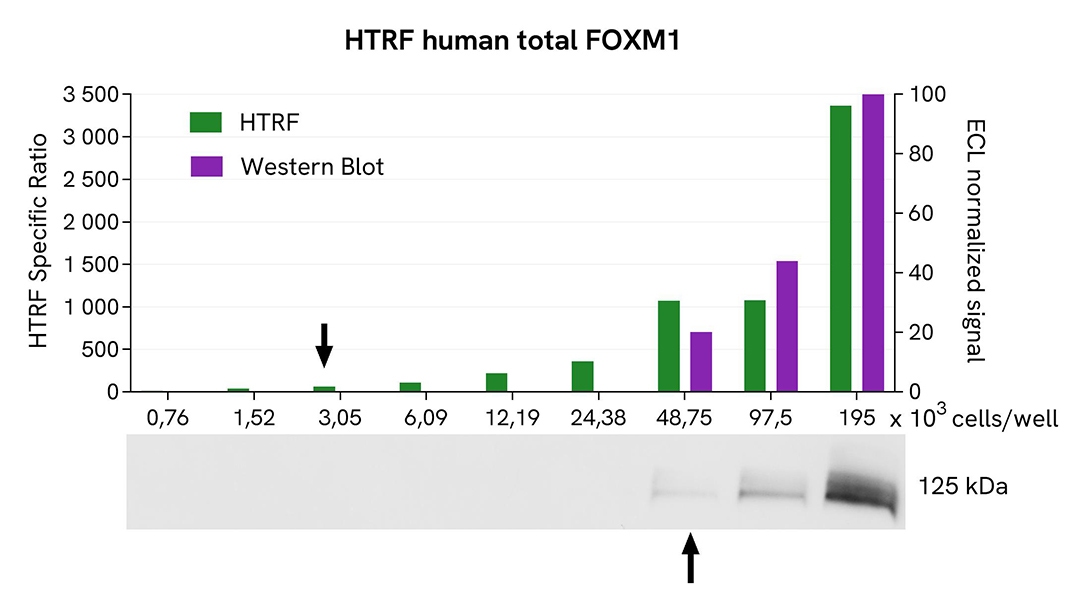
HTRF Total FOXM1 assay compared to Western Blot
HeLa cells were grown in a T175 flask in complete culture medium at 37°C, 5% CO2, until 80% confluence. After a 24h incubation, the cells were lysed with 3 mL of supplemented lysis buffer #1 (1X) for 30 minutes at RT under gentle shaking. Serial dilutions of the cell lysate were performed using supplemented lysis buffer, and 16 µL of each dilution were transferred into a low volume white microplate before the addition of 4 µL of HTRF Total FOXM1 detection reagents. Equal amounts of lysates were used for a side-by-side comparison between HTRF and Western Blot. In these conditions, the HTRF Total FOXM1 assay was 16 times more sensitive than the Western Blot technique.
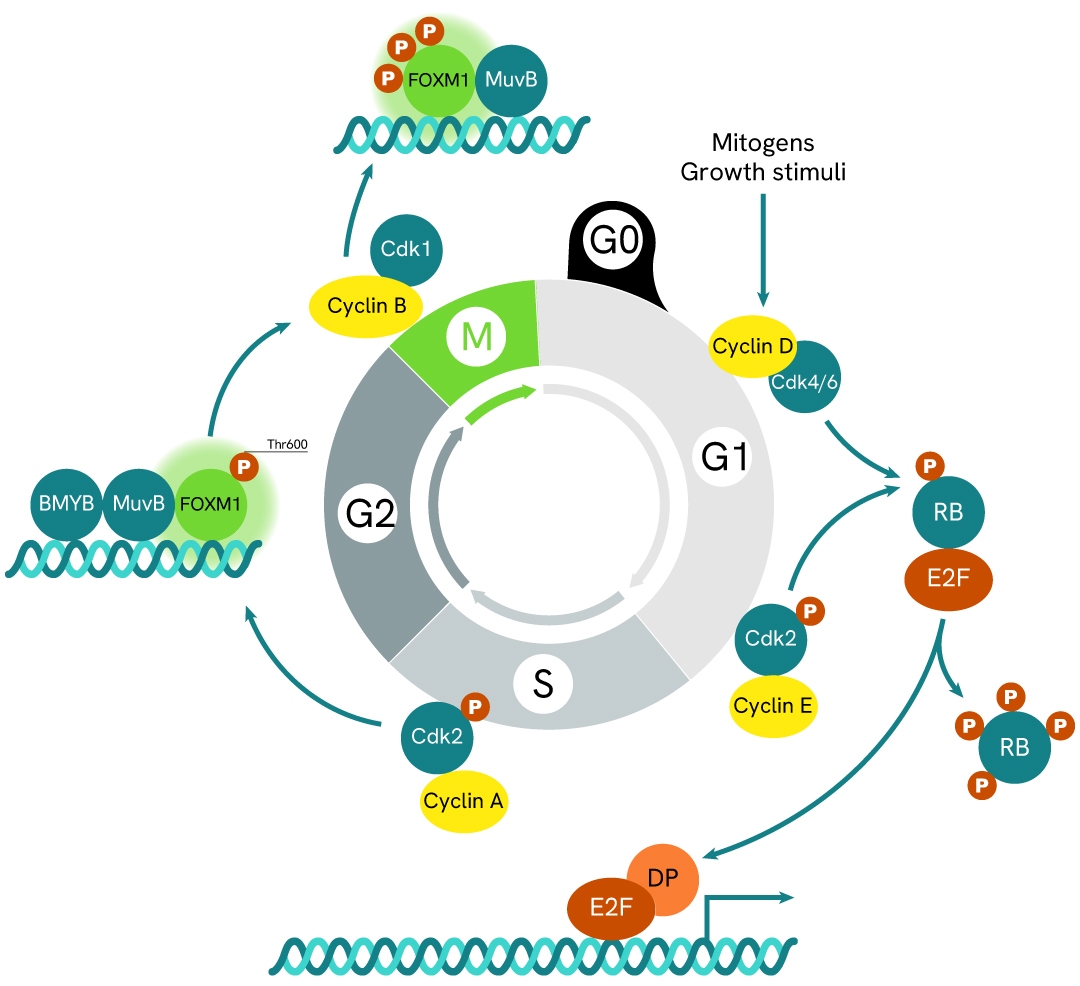
Simplified pathway
FOXM1 signaling pathway
FOXM1 is a regulatory factor involved in the G1/S and G2/M phases of the cell cycle. Its activity is controlled by multisite phosphorylation and ubiquitination throughout the cell cycle. The transcriptional activity of FOXM1 is tightly regulated by phosphorylation at multiple sites by various kinases and phosphatases, as well as by proteasomal degradation. During the S phase, FOXM1 is phosphorylated by cyclin E/A–CDK2 complexes, followed by hyperphosphorylation in the G2 and M phases by cyclin B–CDK1. Several genes and transcription factors, including p53, WDR5, and HIF‑1α, regulate FOXM1 expression, contributing to increased cell invasion, epithelial–mesenchymal transition (EMT), inhibition of apoptosis, and enhanced cell survival.
Specifications
| Application |
Cell Signaling
|
|---|---|
| Brand |
HTRF
|
| Detection Modality |
HTRF
|
| Lysis Buffer Compatibility |
Lysis Buffer 1
Lysis Buffer 2
Lysis Buffer 3
Lysis Buffer 4
|
| Molecular Modification |
Total
|
| Product Group |
Kit
|
| Sample Volume |
16 µL
|
| Shipping Conditions |
Shipped in Dry Ice
|
| Target |
FOXM1
|
| Target Class |
Phosphoproteins
|
| Target Species |
Human
|
| Technology |
TR-FRET
|
| Therapeutic Area |
Oncology
|
| Unit Size |
10,000 assay points
|
Resources
Are you looking for resources, click on the resource type to explore further.
Loading...


How can we help you?
We are here to answer your questions.






























