
HTRF Cellular GTP / Gi protein Binding Kit, 500 assay points
The HTRF Cellular GTP / Gi protein Binding Kit is an immunoassay designed for the detection and quantitation of GTP / Gi protein binding as a readout of Gi-coupled GPCR activity in cells, in a homogeneous (no wash steps, no separation steps) format.
| Feature | Specification |
|---|---|
| Application | Second Messenger Detection |
The HTRF Cellular GTP / Gi protein Binding Kit is an immunoassay designed for the detection and quantitation of GTP / Gi protein binding as a readout of Gi-coupled GPCR activity in cells, in a homogeneous (no wash steps, no separation steps) format.


Loading...
Product information
Overview
This cellular GTP / Gi binding assay is a fluorescent and homogeneous format to measure Gi protein activity in cells at the level of the GDP/GTP nucleotide exchange that characterizes GPCR activation. As such, it has the advantage of monitoring the activation of GPCRs at the level of one of the earliest receptor-mediated events. It therefore provides insight into the upstream portion of GPCR signaling, useful when studying functional responses.
HTRF assays offer many advantages over other technologies:
- Homogeneous add-and-read format
- No wash steps
- Low background
- Straightforward miniaturization from 96- or 384-well microplates to high density assay formats such as 384-well low volume and 1536-well plates
- Stable signal, providing flexibility in readout time or size of assays
How it works
GTP Gi cellular binding assay principle
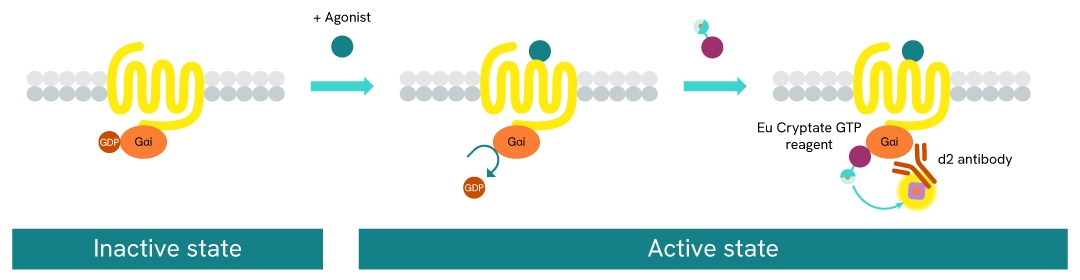
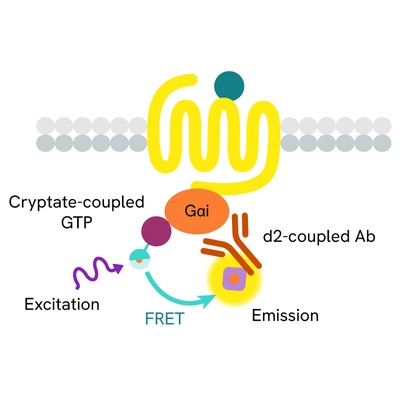
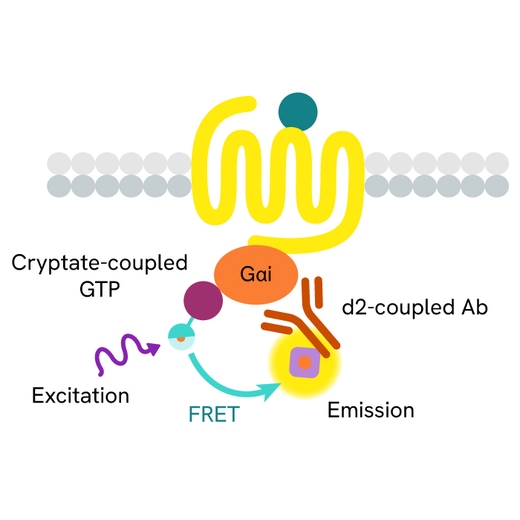
The GTP/Gi cellular binding assay is intended for the simple, rapid, and direct detection of Gαi protein activation in permeabilized cells. It is an upstream readout of Gi protein coupled receptor activation. GPCR activation leads to GDP/GTP nucleotide exchange into the Gα subunit. The principle of this assay is based on HTRF™ technology. It uses a non-hydrolysable GTP analog coupled to the fluorescent Europium cryptate donor.
In practice, agonist-induced GPCR stimulation leads to Gα protein conformation change, and the replacement of Gα-bound GDP by the fluorescent GTP analog in the corresponding binding pocket. Detection is made possible by the addition of d2-labeled anti-Gαi monoclonal antibody (red acceptor). When Europium cryptate and d2 are brought into close proximity, the time-resolved energy transfer between them triggers a TR-FRET signal at d2. This specific signal is proportional to the Gαi activation state. The assay enables the direct pharmacological characterization of compounds acting on Gαi-coupled receptors in permeabilized cells.

GTP Gi cellular binding assay protocol
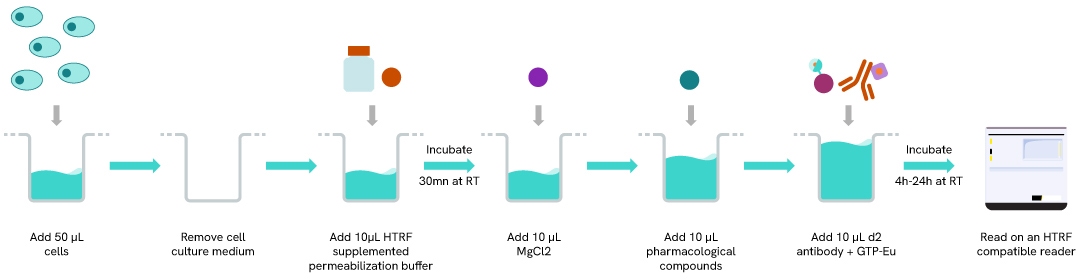
The GTP/Gi cellular binding assay is performed in a miniaturized format using 40 µl final volume and features a streamlined 4-step reagent addition protocol where permeabilization, stimulation, and detection steps are performed successively after plating cells for 24h. Plates are read after 4H or ON incubation at RT, depending on the biological model.
Optimal cell densities, MgCl2 concentrations, and incubation times before reading must be determined based on the chosen biological model (cellular model/receptor/pharmacological compound).
Assay validation
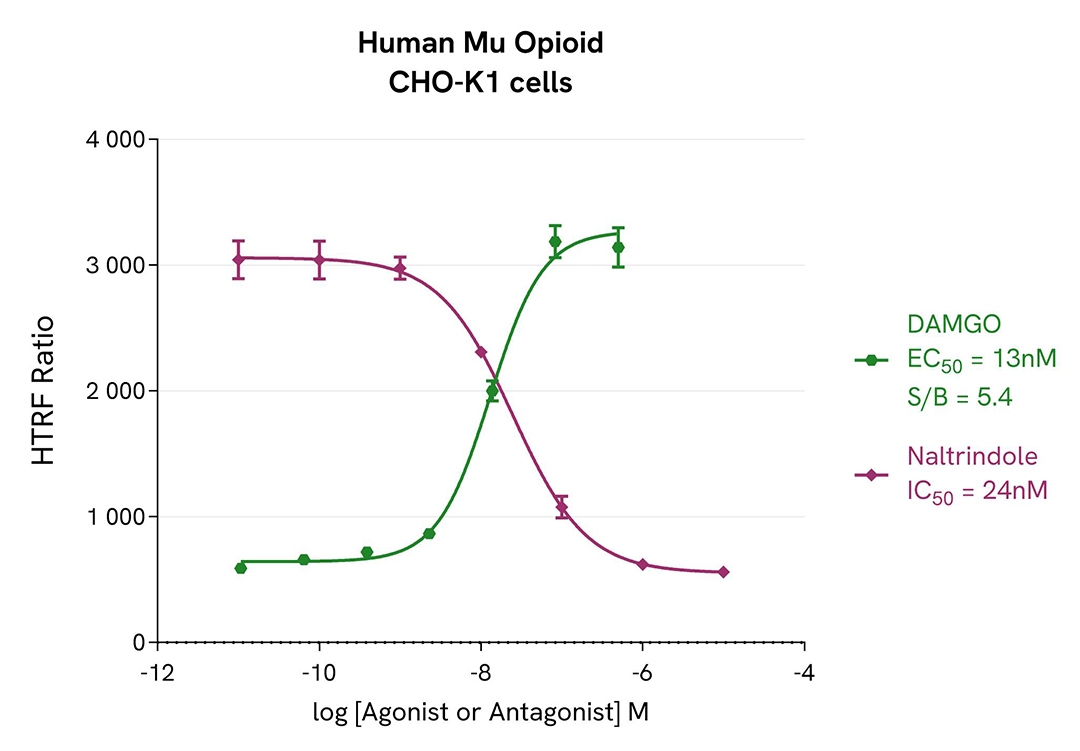
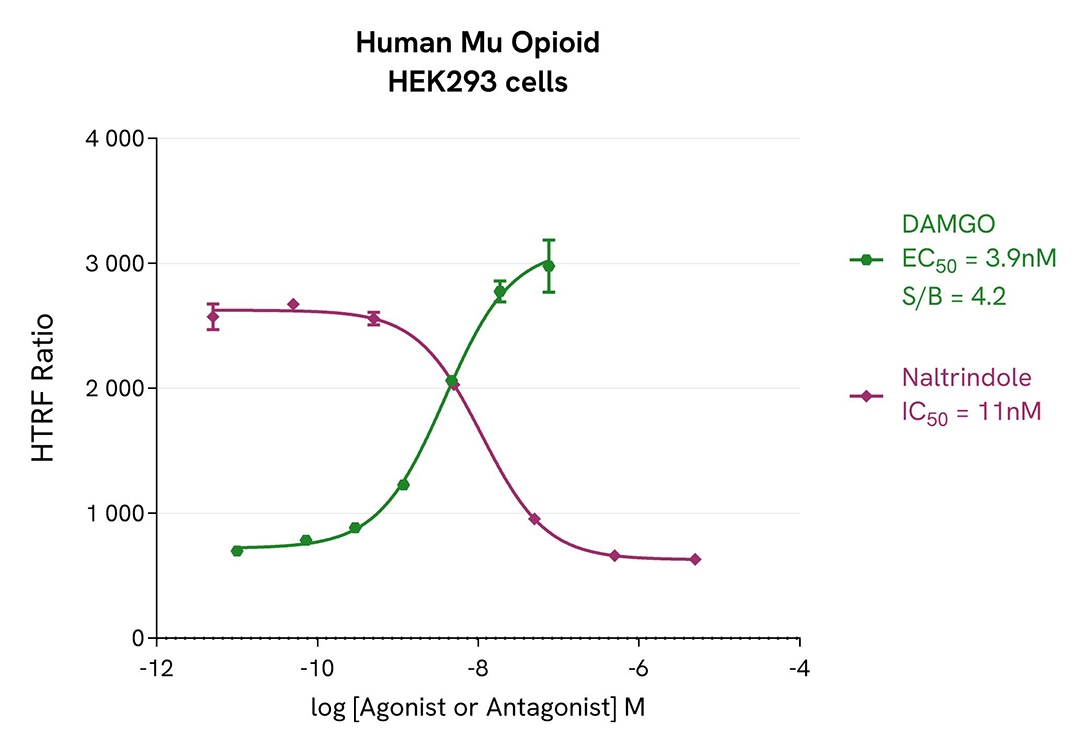
Agonist and antagonist dose-response curves in CHO-K1 & HEK 293 Human Mu opioid (MOR) cell lines
The assay was performed using previously optimized conditions for the chosen biological model and following the assay protocol provided in the kit manual. In particular, the optimal MgCl2 concentration for the stimulation step in these conditions was determined to be 25 mM.
For the stimulation step, Tag-lite CHO-K1 Human Mu opioid (MOR) cells (Revvity #C2SU1MOP) and Tag-lite HEK293 Human Mu opioid cells were incubated overnight in presence of Mu opioid agonist (DAMGO) or antagonist (Naltrindole).
The results were consistent with published data. After incubation, Naltrindole and DAMGO respectively exhibited full antagonist and agonist profiles in both CHO-K1 and HEK 293 cells. The assay S/B is satisfactory in both cellular models, and we report EC50 & IC50 values in line with the literature.
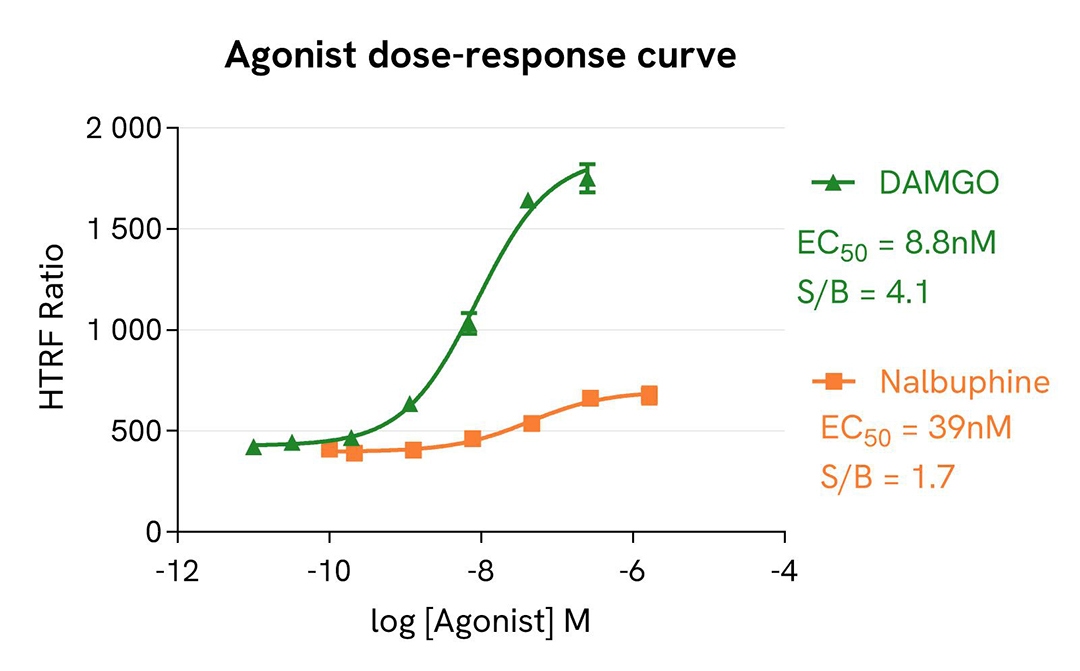
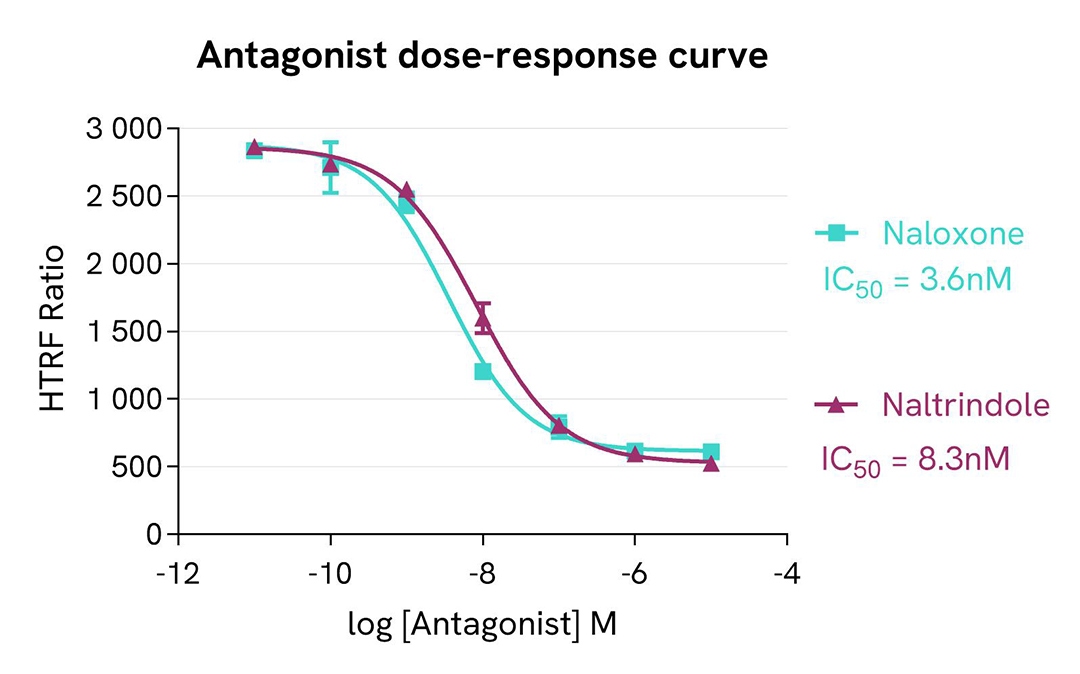
Pharmacological screening in CHO-K1 Human Mu opioid (MOR) cell line
The assay was performed using previously optimized conditions for the chosen biological model and following the assay protocol provided in the kit manual. In particular, the optimal MgCl2 concentration for the stimulation step in these conditions was determined to be 25 mM.
For the stimulation step, Tag-lite CHO-K1 Human Mu opioid (MOR) cells (Revvity #C2SU1MOP) were incubated overnight with MOR agonists (DAMGO & Nalbuphine) and antagonists (Naltrindole & Naloxone).
The results were consistent with published data. After incubation, Naltrindole and Naloxone exhibited full antagonist profiles. DAMGO and Nalbuphine respectively present full agonist and partial agonist profiles. The EC50 & IC50 values and pharmacological rankings derived from these are in line with values reported in the literature.
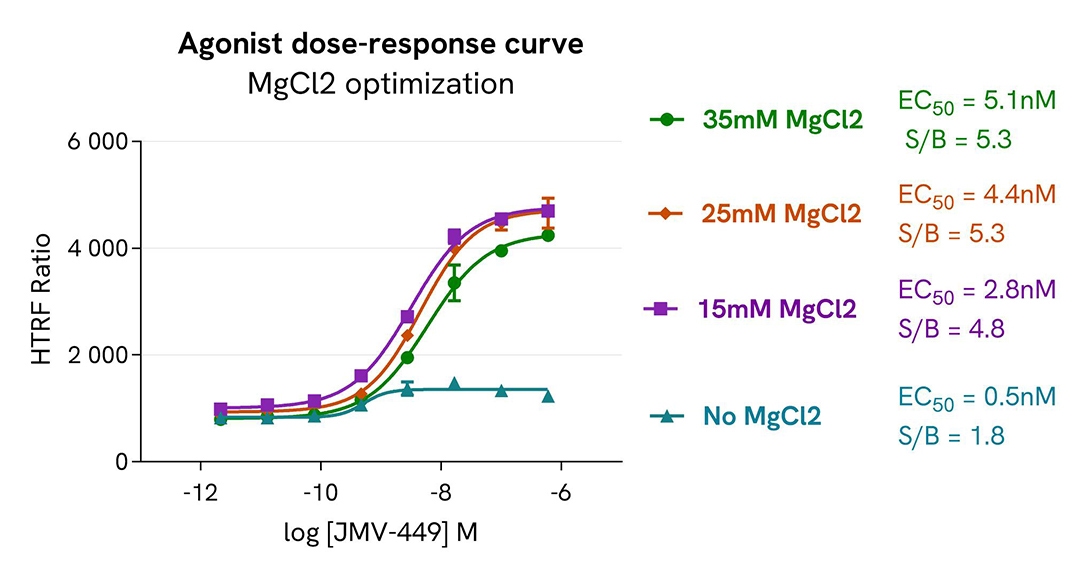
MgCl2 optimization in CHO-K1 Human Neurotensin (NST1) cell line
The assay was performed following the protocol provided in the kit manual. Prior to the experiment, the ValiScreen CHO-K1 Human Neurotensin (NST1) cell line (Revvity #ES-690-C) was plated and cultivated for 24h in suitable conditions.
For the stimulation step, the cells were incubated with increasing concentrations of NTS1 receptor agonist JMV-449, in the absence of MgCl2 or in the presence of MgCl2 concentrations ranging from 15mM to 35mM.
The results were consistent with published data. JMV-449 dose-response curves show a full agonist profile under all conditions. As expected, the MgCl2 improved the signal-to-background (S/B) ratio but produced a slightly detrimental effect on the EC₅₀ value in this biological model.
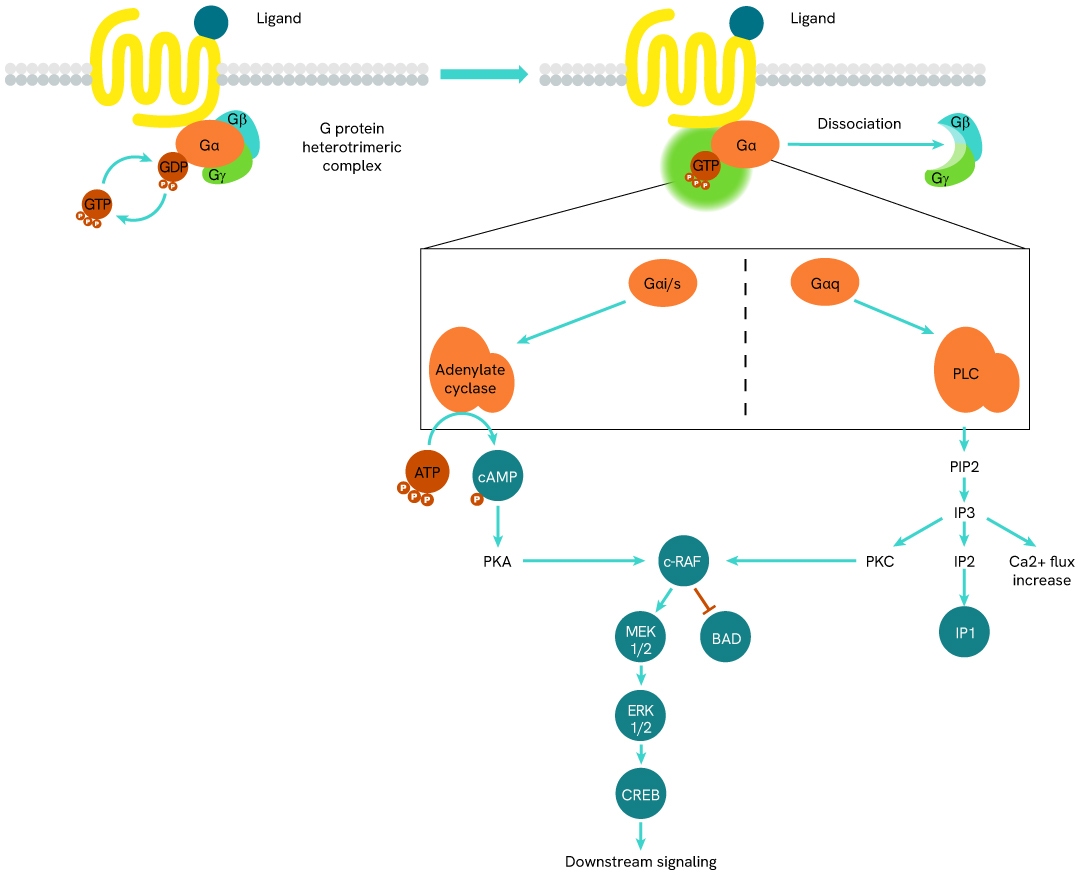
Simplified pathway
GTP role in GPCR signalling pathway
In their inactive state, heterotrimeric G proteins bind GDP in a binding pocket of their Ga sub-unit. Upon stimulation of the GPCR by an agonist, the receptor-coupled G protein undergoes a conformational change that allows the Ga sub-unit to release its bound GDP and replace it with GTP. This nucleotide exchange fully activates the Ga sub-unit, which dissociates from the Gb/Gg dimer and performs inhibitory or stimulating functions on adenylate cyclase (AC) and phospholipase C (PLC), depending on its sub-type (Gai, Gas, Gaq, etc).
The following signal transduction steps are GPCR-specific and mobilize different phospho-proteins and pathways. Over time, the Ga-bound GTP undergoes hydrolysis, which returns the Ga protein to an inactive state.
Specifications
| Application |
Second Messenger Detection
|
|---|---|
| Brand |
HTRF
|
| Detection Modality |
HTRF
|
| Product Group |
Kit
|
| Shipping Conditions |
Shipped in Dry Ice
|
| Target |
Gi protein
|
| Target Class |
GPCR
|
| Technology |
TR-FRET
|
| Therapeutic Area |
Cardiovascular
Infectious Diseases
Inflammation
Metabolism/Diabetes
NASH/Fibrosis
Neuroscience
Oncology & Inflammation
Rare Diseases
|
| Unit Size |
500 assay points
|
Resources
Are you looking for resources, click on the resource type to explore further.
Loading...


How can we help you?
We are here to answer your questions.






























