
RiboLace Pro


RiboLace® Pro is an affinity-based capture system for ribosome profiling (Ribo-seq) and translatome analysis. It uses a proprietary biotinylated puromycin derivative (3P) to selectively bind translation-engaged ribosomes, enabling magnetic enrichment of ribosome-protected fragments (RPFs) for sequencing. This targeted enrichment focuses footprint libraries on actively translated transcripts, reducing background from inactive or non-translating ribosomal complexes.
RiboLace Pro is suitable for eukaryotic cell lines and tissue samples, and the resulting ribosome-protected fragments are compatible with common RPF library preparation methods used for Ribo-seq. Revvity has validated compatibility with the NEXTFLEX™ Small RNA-Seq Kit. The enrichment step can also be integrated with other small RNA-style library preparation approaches.
RiboLace® Pro is an affinity-based capture system for ribosome profiling (Ribo-seq) and translatome analysis. It uses a proprietary biotinylated puromycin derivative (3P) to selectively bind translation-engaged ribosomes, enabling magnetic enrichment of ribosome-protected fragments (RPFs) for sequencing. This targeted enrichment focuses footprint libraries on actively translated transcripts, reducing background from inactive or non-translating ribosomal complexes.
RiboLace Pro is suitable for eukaryotic cell lines and tissue samples, and the resulting ribosome-protected fragments are compatible with common RPF library preparation methods used for Ribo-seq. Revvity has validated compatibility with the NEXTFLEX™ Small RNA-Seq Kit. The enrichment step can also be integrated with other small RNA-style library preparation approaches.


Loading...
Product information
Overview
RiboLace Pro enables selective enrichment of actively translating ribosomes for ribosome profiling (Ribo-seq) experiments. The technology uses a proprietary puromycin derivative (3P) coupled to magnetic beads to capture translation-engaged ribosomal complexes and the ribosome-protected fragments (RPFs) they contain following nuclease digestion. The antibody-free, tag-free enrichment workflow provides an alternative to gradient ultracentrifugation-based approaches and supports Ribo-seq sample preparation from primary or immortalized eukaryotic cell lines (fresh or frozen) as well as tissue samples. The recovered footprints are compatible with standard RPF library preparation workflows. Compatibility with the NEXTFLEX Small RNA-Seq Kit has been shown while remaining compatible with other common Ribo-seq library preparation approaches.
Key features:
- Selective capture of actively translating ribosomes using 3P-functionalized magnetic beads
- Antibody-free, tag-free enrichment designed to avoid ultracentrifugation-based workflows
- Supports primary or immortalized eukaryotic cell lines (fresh or frozen) and tissue samples
- Enriches ribosome-protected fragments (RPFs) suitable for ribosome profiling and footprint sequencing
- Validated compatibility with the NEXTFLEX Small RNA-Seq Kit
Additional product information
Mechanism of active ribosome capture in Ribo-seq workflows
RiboLace Pro exploits the interaction between a puromycin analog (3P) and the ribosomal A-site to selectively capture actively translating ribosomes. Puromycin mimics the 3′ end of aminoacyl-tRNA and can enter the catalytic center of ribosomes engaged in peptide elongation. In RiboLace Pro, the molecule is immobilized on magnetic beads, allowing translation-competent ribosomal complexes to be isolated through affinity capture.
Because the interaction relies on puromycin-like chemistry associated with the catalytic center of active ribosomes, the enrichment step preferentially isolates ribosomes engaged in translation relative to inactive or dissociated complexes. Following nuclease digestion, these ribosomes remain associated with short RNA fragments that correspond to ribosome-protected regions of transcripts.
Magnetic separation of the captured complexes therefore enriches ribosome-protected fragments derived from actively translated mRNA regions. These fragments can then be purified and processed using standard Ribo-seq library preparation workflows to measure ribosome occupancy and translation dynamics across the transcriptome.
Advantages over sucrose gradients and polysome profiling
Many ribosome profiling workflows use sucrose gradient fractionation or polysome profiling to separate translating ribosomes as part of upstream enrichment. These approaches rely on ultracentrifugation and fraction collection, which can add complexity and require access to an ultracentrifuge.
RiboLace Pro simplifies this step by enriching translating ribosomes directly from cell lysates using an affinity-based pull-down. Because the enrichment occurs without gradient separation, the workflow can be performed without fractionation or ultracentrifugation while still producing material suitable for downstream Ribo-seq analysis.
Library preparation and QC tools for Ribo-seq workflows
RiboLace Pro enrichment workflows integrate with standard small RNA-style library preparation approaches used for ribosome profiling. Compatibility with the NEXTFLEX Small RNA-Seq Kit is demonstrated in our application note, providing a clear path from enriched ribosome footprints to sequencing-ready libraries using a workflow optimized for short RNA inserts.

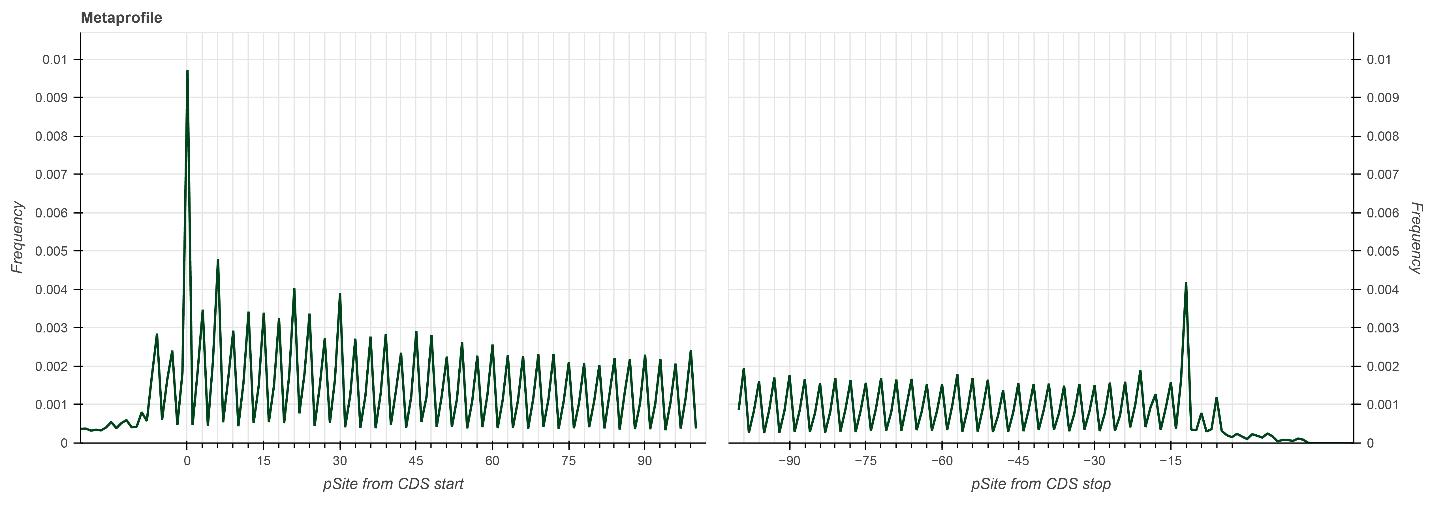
Figure 1. Metaprofile plot showing P-site positions for all transcripts around translation initiation sites (0 to +100 nt) and translation termination sites (0 to −100 nt). RPFs were isolated using RiboLace® Pro, and sequencing libraries were prepared using the NEXTFLEX Small RNA-Seq Kit.
Additional tools within the NEXTFLEX ecosystem can support library quality and experimental consistency. miND™ spike-in controls can be incorporated to monitor technical variation and assist with normalization across samples. Custom small RNA blockers can be used as an optional tool to suppress abundant non-target short fragments when they impact usable sequencing depth.
Specifications
| Product Group |
Small RNA Accessory
|
|---|---|
| Shipping Conditions |
Dual Temperature
|
| Unit Size |
12 rxns
|
FAQs
-
What does RiboLace® Pro capture in a ribosome profiling experiment?
-
What types of samples can be used with RiboLace Pro?
-
How does RiboLace Pro differ from sucrose gradients or polysome profiling?
-
What library preparation workflows are compatible with RiboLace Pro?
-
Does RiboLace Pro capture stalled ribosomes?
-
What sequencing depth is recommended for Ribo-seq experiments?
Resources
Are you looking for resources, click on the resource type to explore further.
Flyer illustrating the utility of RiboLace Pro in active ribosome profiling.
This flyer shows Revvity's library prep & accessory solutions for Small RNA-seq.
Application note illustrating the compatability of RiboLace & NEXTFLEX Small RNA v4.
Loading...


How can we help you?
We are here to answer your questions.






























