
MERCURIUS Total Drug-seq kits





MERCURIUS™ Total DRUG-seq is an extraction-free total RNA-seq solution for full-length transcriptome profiling at screening scale. Designed to work directly from cell lysates, it brings broader RNA coverage into large chemical and genetic perturbation studies, enabling analysis of coding and non-coding RNAs, differential gene expression, transcript variants, alternative splicing, fusion genes, and promoter usage.
For researchers who need more than a 3′ readout, MERCURIUS Total DRUG-seq combines multiplexed library preparation with a one-day workflow to support target discovery, target validation, and deeper transcriptomic analysis without prior RNA extraction. Compatible with Illumina® and AVITI™ sequencers, it offers a scalable route to full-length RNA insight in screening and discovery workflows.
| Feature | Specification |
|---|---|
| Barcodes | 4 UDI Pairs Included |
MERCURIUS™ Total DRUG-seq is an extraction-free total RNA-seq solution for full-length transcriptome profiling at screening scale. Designed to work directly from cell lysates, it brings broader RNA coverage into large chemical and genetic perturbation studies, enabling analysis of coding and non-coding RNAs, differential gene expression, transcript variants, alternative splicing, fusion genes, and promoter usage.
For researchers who need more than a 3′ readout, MERCURIUS Total DRUG-seq combines multiplexed library preparation with a one-day workflow to support target discovery, target validation, and deeper transcriptomic analysis without prior RNA extraction. Compatible with Illumina® and AVITI™ sequencers, it offers a scalable route to full-length RNA insight in screening and discovery workflows.






Loading...
Product information
Overview
Not every screening study can be answered with a 3′ counting assay. When the biology depends on transcript structure, non-coding RNA, or promoter switching, MERCURIUS Total DRUG-seq brings full-length transcriptome information into chemical and genetic perturbation screening workflows in an extraction-free format designed for scale. Working directly from cell lysates, it supports multiplexed total RNA library preparation for large compound and genetic perturbation studies, target validation, and toxicology studies.
That added depth matters when differential expression alone is not enough. MERCURIUS Total DRUG-seq is positioned for analysis of coding and non-coding RNAs, transcript variants, alternative splicing, fusion genes, and promoter usage, while delivering reads across the full gene body rather than a 3′-biased view. With a one-day workflow and compatibility with Illumina and AVITI sequencers, it gives researchers a more practical route to deeper transcriptomic analysis at screening scale.
Key features:
- Extraction-free total RNA-seq workflow from cell lysates
- Multiplexed full-length RNA workflow with 96- or 384-sample pooling
- Broader RNA coverage across coding and non-coding transcripts
- Supports analysis of transcript variants, alternative splicing, fusion genes, and promoter usage
- One-day workflow compatible with Illumina and AVITI sequencing platforms
Additional product information
When a 3′ readout is not enough
Some discovery questions need more than gene-level counting. When the biology depends on transcript structure, non-coding RNA, or events across the full gene body, a 3′-biased assay can leave important biology unresolved. MERCURIUS™ Total DRUG-seq is positioned for those studies, combining extraction-free total RNA-seq with full-length transcriptome profiling in a format built for chemical and genetic perturbation screening and target discovery.

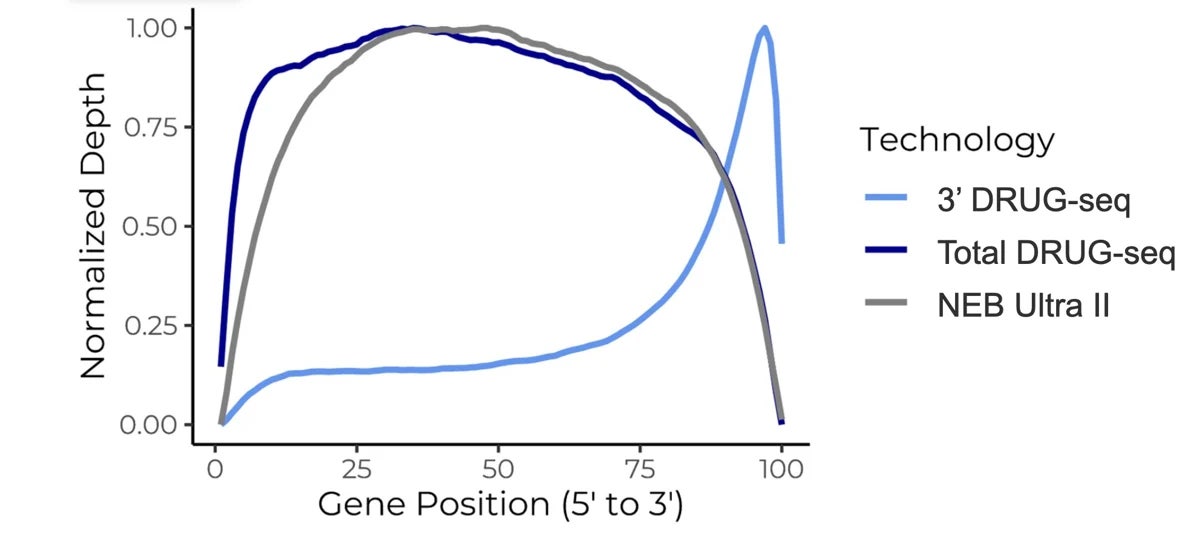
Figure 1. Total DRUG-seq provides read coverage across the full gene body, supporting transcriptome analysis beyond the 3′-biased view of standard 3′ DRUG-seq.
Working directly from cell lysates, the assay brings broader RNA information into large-scale workflows without a separate RNA extraction step. That makes it a strong fit when a 3′ snapshot is no longer enough, but screening-scale practicality still matters.
Go beyond differential expression with transcript variants, splicing, and fusion genes
Differential expression can show that transcription changed. It does not always show how. MERCURIUS™ Total DRUG-seq is positioned for applications that require more transcript-level detail, including transcript variants, alternative splicing, fusion genes, and promoter usage.
That added resolution matters when transcript architecture is part of the biological question, not just overall gene abundance. For studies that need to interpret complex responses rather than simple gene-count changes, full-length transcript coverage can provide a more informative view of what changed across the transcriptome.
Full-length transcriptome profiling for target discovery and validation
Target discovery and target validation often require more than a high-level expression signature. Broader RNA coverage can add clarity when the goal is to understand pathway engagement, transcript complexity, or downstream consequences of perturbation in greater detail. MERCURIUS™ Total DRUG-seq is positioned for full-length transcriptomic profiling in chemical and genetic perturbation workflows, target discovery, and target validation.
The workflow is also designed to remain usable at scale. The user guide describes kit formats with 96- or 384-sample multiplexing depending on configuration, allowing samples to be pooled in a streamlined multiplexed workflow.
A natural fit for Revvity discovery workflows
MERCURIUS Total DRUG-seq fits naturally into Revvity discovery workflows and can be automated on the Fontus™ workstation for higher-throughput implementation. It complements PhenoVue™ Cell Painting Kits and high-content imaging systems such as Opera Phenix™ Plus and Operetta® CLS™ by adding full-length transcriptome information when transcript structure, pathway-level follow-up, or target hypotheses require more than a phenotypic or 3′ readout. Together with Revvity’s Dharmacon™ RNAi and CRISPR screening tools, these solutions can help address questions related to genes, pathways, and phenotypes in drug discovery.
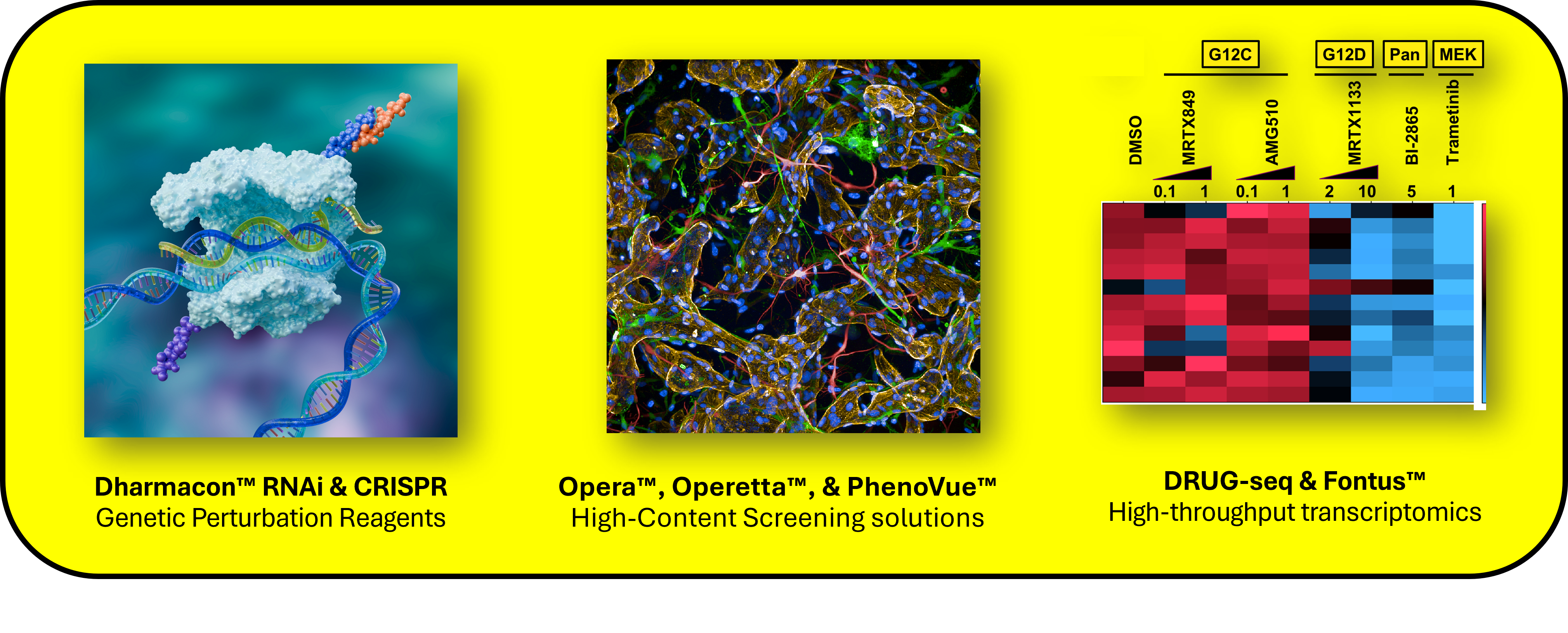
Figure 2: Revvity solutions for drug discovery, including Dharmacon™ genetic perturbation tools, Opera™ and Operetta™ high-content imaging with PhenoVue™, and MERCURIUS™ Total DRUG-seq with Fontus™ for transcriptional profiling. Together, these help researchers address questions related to genes, pathways, and phenotypes at screening scale.
Specifications
| Barcodes |
4 UDI Pairs Included
|
|---|---|
| Format |
Sample Multiplexing: 1 x 384 well-plate
|
| Product Group |
RNA Sequencing
|
| Shipping Conditions |
Dual Temperature
|
| Unit Size |
384 preps
|
Video gallery
FAQs
-
When should I choose MERCURIUS™ Total DRUG-seq instead of standard DRUG-seq?
-
What sequencing platforms and kit formats are supported?
-
What sample input does the workflow require?
-
Can Total DRUG-seq support transcript-level as well as gene-level analysis?
-
Which species and sample types are supported?
Resources
Are you looking for resources, click on the resource type to explore further.
Loading...


How can we help you?
We are here to answer your questions.






























