
MERCURIUS Drug-seq kits



MERCURIUS™ DRUG-seq is a high-throughput transcriptomics solution for screening-scale gene expression profiling, using an extraction-free 3′ mRNA-seq workflow to turn cell lysates directly into sequencing-ready libraries. Designed for 96-, 384-, and 1,536-well formats and enabled by early multiplexing, it reduces the time, cost, and workflow burden of conventional RNA-seq while delivering gene expression signatures at screening scale for drug-target and functional genomics workflows, hit prioritization, and mechanism-of-action studies.
When phenotypic screening shows what changed but not why, DRUG-seq adds pathway-level insight by revealing transcriptional responses across thousands of perturbations. Researchers can use those signatures to cluster compounds by biological response, investigate potential off-target activity, compare genetic perturbations by transcriptional effect, and connect imaging phenotypes to underlying mechanism. Within the Revvity screening workflow, DRUG-seq complements high-content imaging, Cell painting and Dharmacon™ RNAi & CRISPR screening by adding transcriptomic context for higher-confidence MoA profiling.
| Feature | Specification |
|---|---|
| Automation Compatible | Yes |
| Barcodes | 4 UDI Pairs Included |
MERCURIUS™ DRUG-seq is a high-throughput transcriptomics solution for screening-scale gene expression profiling, using an extraction-free 3′ mRNA-seq workflow to turn cell lysates directly into sequencing-ready libraries. Designed for 96-, 384-, and 1,536-well formats and enabled by early multiplexing, it reduces the time, cost, and workflow burden of conventional RNA-seq while delivering gene expression signatures at screening scale for drug-target and functional genomics workflows, hit prioritization, and mechanism-of-action studies.
When phenotypic screening shows what changed but not why, DRUG-seq adds pathway-level insight by revealing transcriptional responses across thousands of perturbations. Researchers can use those signatures to cluster compounds by biological response, investigate potential off-target activity, compare genetic perturbations by transcriptional effect, and connect imaging phenotypes to underlying mechanism. Within the Revvity screening workflow, DRUG-seq complements high-content imaging, Cell painting and Dharmacon™ RNAi & CRISPR screening by adding transcriptomic context for higher-confidence MoA profiling.



Loading...
Product information
Overview
MERCURIUS DRUG-seq is designed for screening-scale transcriptomics where conventional RNA-seq is often too slow and labor-intensive. This extraction-free, plate-based 3′ mRNA-seq workflow converts cell lysates directly into sequencing-ready libraries and enables early sample pooling, helping streamline gene expression profiling across large chemical and genetic perturbation studies.
Rather than deep profiling of a small number of samples, DRUG-seq is built to generate robust gene expression signatures across many conditions. That makes it well suited for drug target and functional genomics workflows, hit prioritization, dose-response studies, and mechanism-of-action work, particularly when paired with phenotypic screening, high-content imaging and Dharmacon RNAi & CRISPR screening.
Key features:
- Extraction-free workflow from cell lysates to reduce hands-on time and simplify library preparation
- Early sample barcoding and pooling to support efficient processing of large screening studies
- 3′ mRNA-seq approach optimized for gene expression signatures across many perturbations
- Compatible with 96-, 384-, and 1,536-well screening formats for scalable transcriptomic profiling workflows
- Supports transcriptional profiling across chemical and genetic perturbation workflows
Specifications
| Automation Compatible |
Yes
|
|---|---|
| Barcodes |
4 UDI Pairs Included
|
| Format |
RNA multiplexing: 384
|
| Product Group |
RNA Sequencing
|
| Shipping Conditions |
Dual Temperature
|
| Unit Size |
384 preps
|
Additional product information
Why standard RNA-seq becomes a bottleneck at screening scale
Transcriptomics can add real value in early discovery, but conventional RNA-seq is often too slow and labor-intensive for screening-scale studies. Once experiments expand to hundreds or thousands of conditions, RNA extraction, individual library prep, and longer turnaround can quickly become limiting.
MERCURIUS™ DRUG-seq is designed to make gene expression profiling more practical at scale. Its extraction-free 3′ mRNA-seq workflow converts cell lysates directly into sequencing-ready libraries and enables early sample pooling, helping streamline transcriptomic profiling across 96-, 384-, and 1,536-well screening formats.
Transcriptomic signatures for mechanism-of-action profiling
Phenotypic assays can show that a compound changes cell state, but they do not always explain how. Likewise, functional genomics screens can identify active genes without fully resolving downstream pathway effects. DRUG-seq adds that missing layer by capturing gene expression responses, giving researchers a way to compare perturbations by biology as well as phenotype.

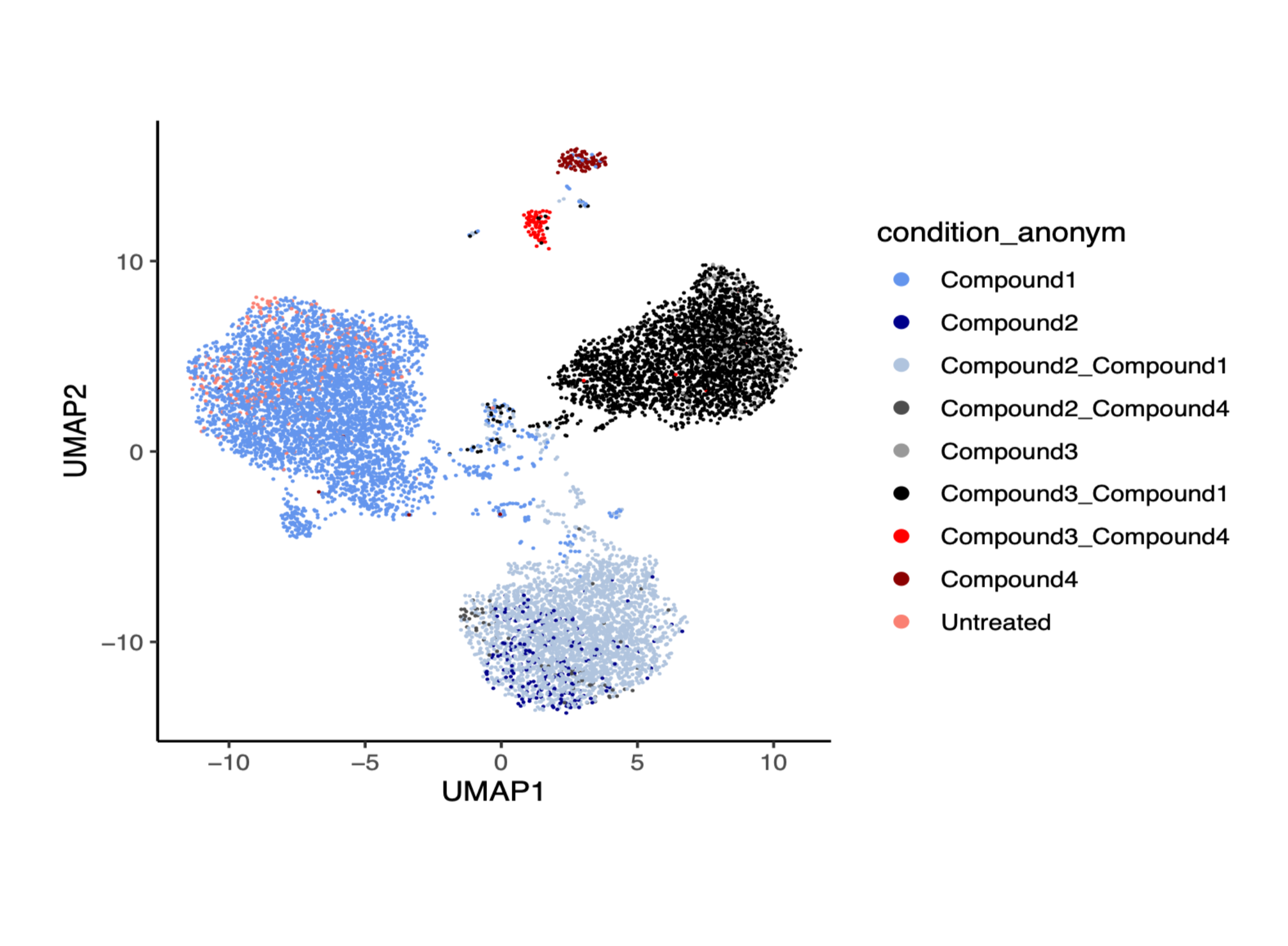
Figure 1. DRUG-seq transcriptomic signatures support clustering of compounds and compound combinations by biological response.
These gene expression signatures can help group compounds with similar mechanisms, differentiate genetic perturbations with shared or distinct transcriptional consequences, and support mechanism-of-action studies earlier in discovery. For teams looking to move beyond surface-level phenotypes, transcriptomic profiling can provide a more informative view of underlying biology.
Better hit triage with molecular context
Hit triage is stronger when it is based on more than visible phenotype alone. Morphology, viability, and other screening readouts can identify active compounds or genetic hits, but they do not always reveal whether those effects reflect the intended biology or a broader, less desirable response.
DRUG-seq helps add that context by generating gene-level signatures across many perturbations in a screening-friendly format. Researchers can use those signatures to compare compound responses, investigate pathway activity, prioritize genes or targets for follow-up, and prioritize hits with greater confidence before moving into more resource-intensive follow-up studies.
A natural fit for Revvity screening workflows
MERCURIUS™ DRUG-seq complements Revvity screening workflows by adding transcriptomic context to phenotypic data and perturbation-driven screening readouts. High-content imaging and Cell Painting can reveal how cells respond to perturbation, while DRUG-seq shows the gene expression changes associated with those phenotypes. In functional genomics workflows, it can also help characterize the biological consequences of genetic perturbations across many conditions and complements Dharmacon™ RNAi & CRISPR screening by adding transcriptional context to genetic perturbation data.
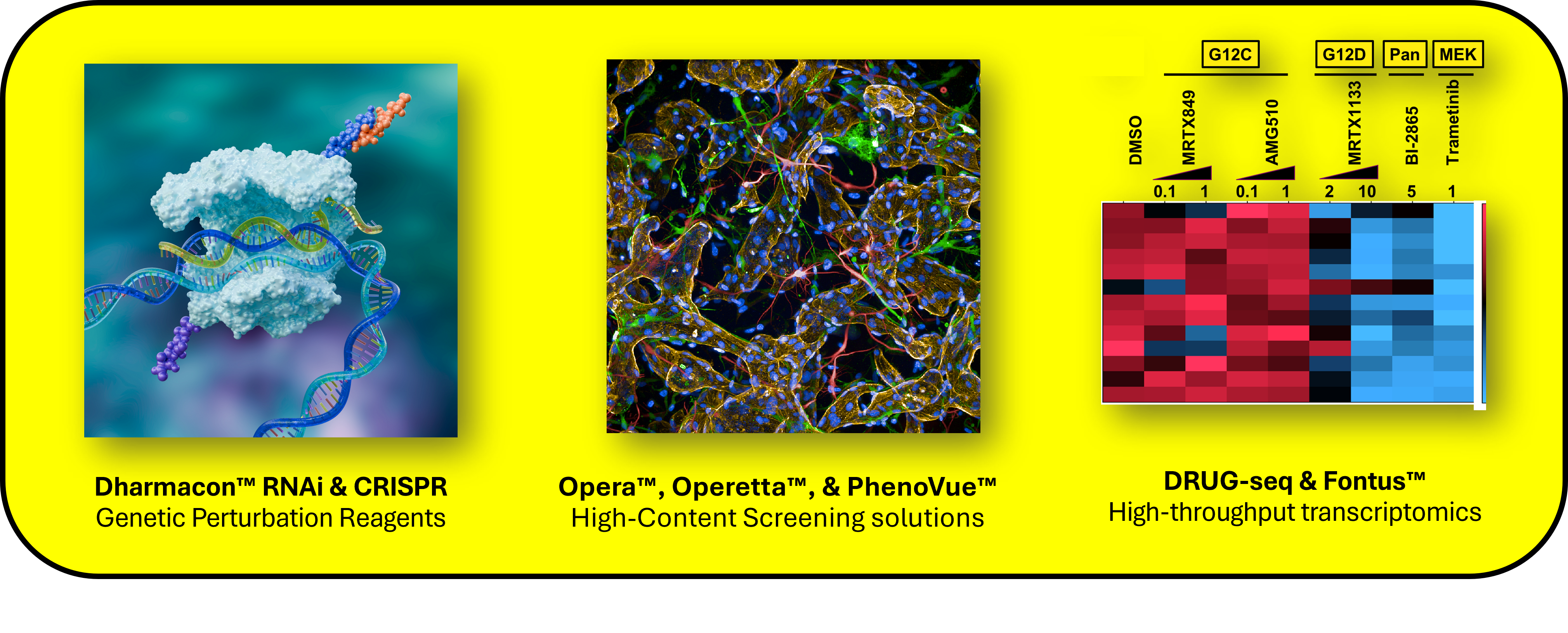
Figure 2: Revvity solutions for drug discovery, including Dharmacon™ genetic perturbation tools, Opera™ and Operetta™ high-content imaging with PhenoVue™, and MERCURIUS™ Total DRUG-seq with Fontus™ for transcriptional profiling. Together, these help researchers address questions related to genes, pathways, and phenotypes at screening scale.
Together, these approaches provide a more complete view of compound, target, or gene-driven biology and support more informed hit prioritization and mechanism-of-action studies.
FAQs
-
When should I choose DRUG-seq instead of Total DRUG-seq?
-
Can DRUG-seq be paired with phenotypic screening or high-content imaging?
-
What sequencing platforms and throughput formats are supported?
-
How much sequencing depth is typically recommended for DRUG-seq libraries?
-
Is DRUG-seq intended for full-length transcript analysis or for screening-scale gene expression profiling?
Resources
Are you looking for resources, click on the resource type to explore further.
Loading...


How can we help you?
We are here to answer your questions.






























